OpenTrons
Earlier this week we started learning about biology lab automation.
There is this YC statup called Opentron which built a robot for automating a large part of biological lab experimental protocols.
An example of a protocol is “Golden gate cloning” which is used to assemble large DNA molecules.
A use case
To frame this in terms of a use case, as I currently understand: say you want to produce something like insulin.
Insulin is a peptide, meaning it is a chain of amino acids. It has a shape in 3D space, which due to chemical bonds etc., naturally fits into the receptor of a cell. When it fits into that receptor, the cell begins to suck in glucose. Insulin is thus what is referred to as a “signalling” molecule. It doesn’t do anything itself. It seems like you can’t get other shapes to fit this receptor.

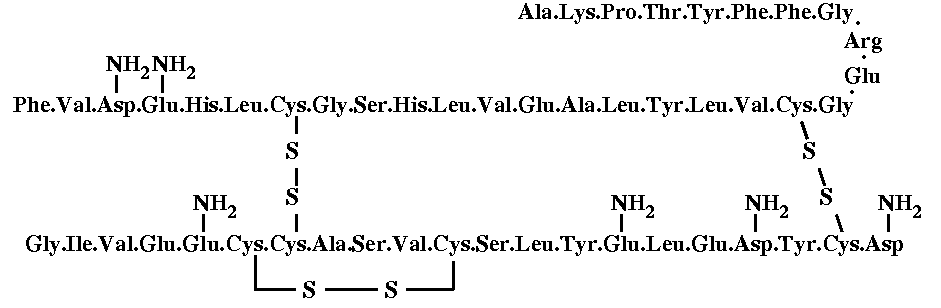
I have diabetes, so I don’t get insulin secreted by my body anymore. So I get it injected extraneously. Insulin is usually made in the pancreas.
Where does it come from?
What is the process of making insulin? How do you make it outside of the body?
When it was first discovered, it was taken from dog organs. They saw some juice excreted from a pancreas and tested its effects.
In 1951, we fully sequenced the amino acid structure of insulin, which is the first step towards making it. What does that mean though?
Biological units like cells, molecules, proteins, are made up of chemical elements. These chemical elements bond to each other in different ways due to physics.
Chemical elements bind together in structures. Some of these structures are categorised as amino acids.
These structures also chain together into longer structures. When amino acids are chained, they are referred to as peptides. A peptide might be 2-50 amino acids chained together. A protein is the term given to larger peptides. Insulin, for example, has a chain of 51 amino acids.
How do we synthesise insulin, a protein (which is a large-ish peptide, which is a chain of amino acids, which is a chain of chemical structures formed together by bonds due to physcs)?
Well, I wish it was as simple as just the compiler to produce the executable, but no. It’s a couple more steps removed. I currently only have a naive understanding of how the body does it.
How a body makes a protein
Here’s how the body produces the protein:
DNA is a string consisting of bases.
type DNA = Base[]
enum Base = {
A, C, G, T
}The human genome is 3 billion base pairs. It is the complete DNA.
let genome: [DNA; 3_000_000_000*2];Every cell in the body contains a copy of the genome, which is the source code for which the entire body is generatively instantiated. There are 30-40 trillion cells in the body.
Same code, different cells
The generative instantiation of an organism from only cells is a dynamic system. The organism proceeds in developmental stages, and when certain thresholds are met, the system transitions into new areas of the landscape.
Each cell develops based on local state. So temperature, light, etc. are some examples of what the cell might sense. A plant seed will sense gravity, and roots will grow downwards into soil. How do roots grow? These are tissues which are composed of differentiated root cells (root epidermal cells). The purpose of roots is to absorb and transport water, for which they have specialised proteins that do this.
All plant cells have the same DNA, but they express different sets of proteins. A root cell will have proteins for water transport. A leaf cell will have proteins for photosynthesis.
How does the organism compute the transition from growing roots to sprouting leaves? The cells in the stem, broaching above ground, will grow to a certain height. At some point, the cells will begin to sense light. Proteins within the cells will broadcast this information to many other cells via the plant’s vascular network. This happens by way of a protein creating a signalling molecule - which we call a hormone. Other cells have receptors which register this type of molecule. When the hormone binds to the cell receptors, it changes the internal dynamics of the cell’s state.
Enough “light” in the “stem” area will shift a region of the plant into producing branches with leaves. Each cell maintains a local state which determines which genes will be read to produce which proteins which produce the form and function of the cell, and this is happening to clusters of millions of cells who all share very similar internal state, which form higher-level features of tissues and organs.



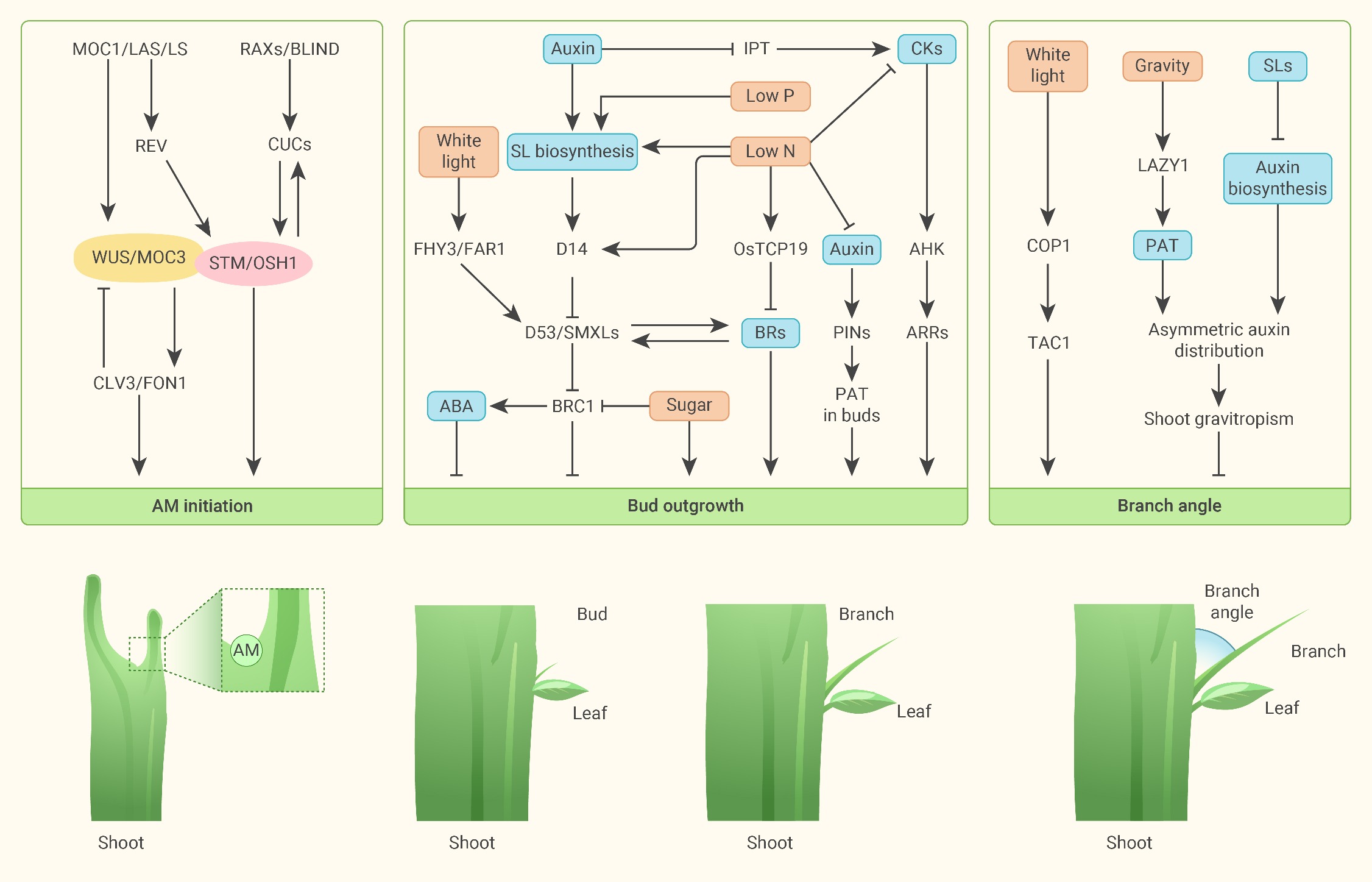
Not a blueprint
This is all from a simple premise that every cell shares the same genome. The genome contains the information which can produce different types of proteins, which change the form and function of a cell, which behave in local networks, which emergently form higher-level structures such as tissues, and this pattern repeats into organs and up until the final organism.
The genome seems like it is closer to neural network weights, rather than a USB of CAD designs + a procedural assembly program.


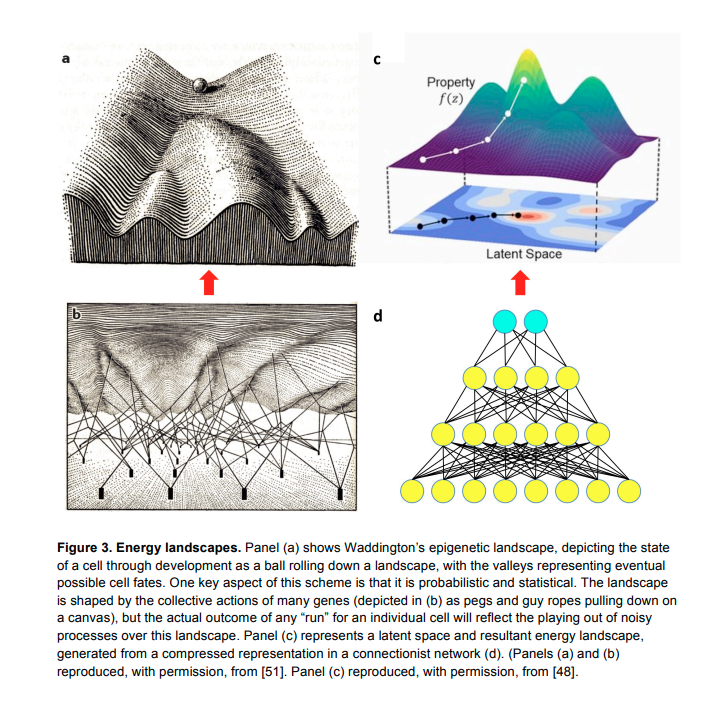
One direction
I got very carried away writing all of this, I was supposed to be talking about how insulin is made. But I find that this mental model is really useful to think through before we actually talk about DNA. Because making proteins is a short-ish pipeline which begins with DNA. Originally I thought the analogy of source code being compiled into lower-level opcodes as a good one, but now I think it’s the wrong shape. Biology resembles more like a generative network than a turing machine.
Insulin is synthesised like any protein is in the body. Here’s how:
- to recap, insulin is defined as a chain of amino acids.
- amino acids chains are assembled step-by-step, this is called translation
- translation: each iteration, we load the current codon from the mRNA, and select the corresponding amino acid to produce, and append to the chain.
- mRNA stores a chain of DNA basis, which begin with a special start token, and a special end token, in order for the translation process to terminate correctly
- mRNA is synthesised by reading areas of the DNA. this is called transcription
- the areas of DNA which are read are determined by a complex dynamics network, specific to the cell, the gene regulatory network (GRN), as well as a few other things (parts of the genome can be frozen, which is passed on to children of that cell)
This process is referred to as the Central Dogma. It’s a one-way (ish) flow of state:

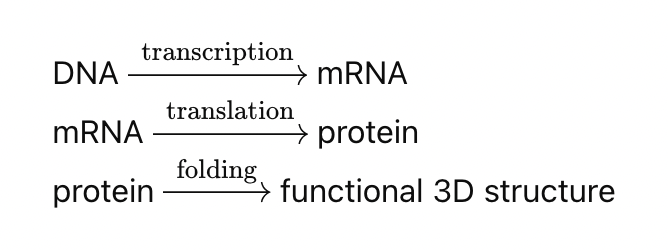
Zooming out
Using bacteria to make it
Now, coming back to the original part of this post - Opentrons, protocols, proteins.
Say you want to produce something like insulin. We can’t print the amino acid chains using a 3D printer. But we can use a couple of ways to create them if we start earlier in the process, from DNA.
One way is by using an existing organism, like bacteria. Bacteria has the same basic machinery of human cells, in that it follows the DNA -> mRNA -> protein pipeline. Unlike human cells, bacterial cultures are significantly cheaper to buy/sell/produce, and are much simpler architecturally, making industrial processes more reliable for the most part.
The way it’s done:
- we take insulin, the protein (a chain of amino acids that folded into a 3d structure)
- we sequence it, to find out its amino acid chain (this occurred in 1951)
- once we know it, we can reverse the amino acids back to the DNA which produces them. This is relatively straightforward, as I understand. It's not reverse-engineering.
- we take the DNA code, and we chemically synthesise it using something like phosphoramidite synthesis. This process has P(success)=(1-0.01)^L, where L is the length of the DNA in bases. Since errors compound exponentially with length, longer DNA is harder to synthesise
- Coming back to the start of this post, in order to synthesise large DNA molecules, we chemically synthesise them in segments, and then join them together using another technique.
- once we have the DNA, we clone it into an expression vector (a plasmid)
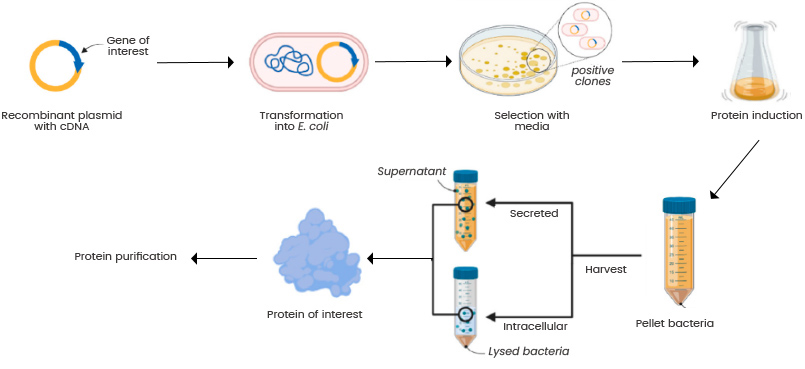
- finally, we are ready to produce it inside an organism like E. Coli bacteria.
- we transform our plasmid into host cells
- we prepare a bench of bacterial cultures. P(yield) is not 1, so we need to replicate this part across many independent bacterial cultures to ensure we have some yield.
- the bacteria is mixed with our plasmid DNA. They are then given a heat shock, which causes some of them to take up the plasmid DNA.
- we detect which cultures took up the DNA (tldr: antibacterial death)
- we pick one colony that took up the DNA, and transfer it into a tube containing the necessary ingredients to grow that colony
- we induce expression of the DNA, by introducing a reagent
- we harvest the liquid culture, which involves purifying and extracting only the proteins
And that’s how insulin is made! It’s called recombinant expression, I believe (again, I am just a student).
Robots in the lab
So how does this relate to lab automation?
Well, earlier on I mentioned that there is this repo of code which runs a specific protocol. A protocol in molecular biology is a concrete sequence of experimental actions a person performs in the lab.
Much like in software and computer programming, people encapsulate complexity in biology too. A protocol is one encapsulation of inputs, process and output.
Unlike software/computer programming, there isn’t really an NPM. There is no package manager to install different protocols - ie. Golden Gate, Gibson assembly.
Opentrons is one sort of project in the direction of that. They provide an API to controlling laboratory robots, which have different primitives, controllers, ingredients, all using Python.
Early days
I am principally interested in biotechnology right now as it is a form of engineering. Compared to software, it is in the early 2000’s of tooling maturity and capability. The engineering infrastructure (cloud labs, programmable lab robotics) seem to be evolving in tandem with scientific discovery (new techniques, new mechanistic models).
This week in the HTGAA course we were learning how to program Opentrons robots. Part of the homework was finding a paper which used Opentrons and describing it. I found one called Slowpoke, which is a protocol generator for doing DNA cloning for large DNA sequences through the Golden Gate cloning technique.
By protocol generator, that means - literally it’s python code to generate more python code. Evidently, we are in the macros era of bio-engineering. :P The reason for this is that Opentrons seem to only accept a single Python executable. It is not a “runtime” yet.
Rewriting it
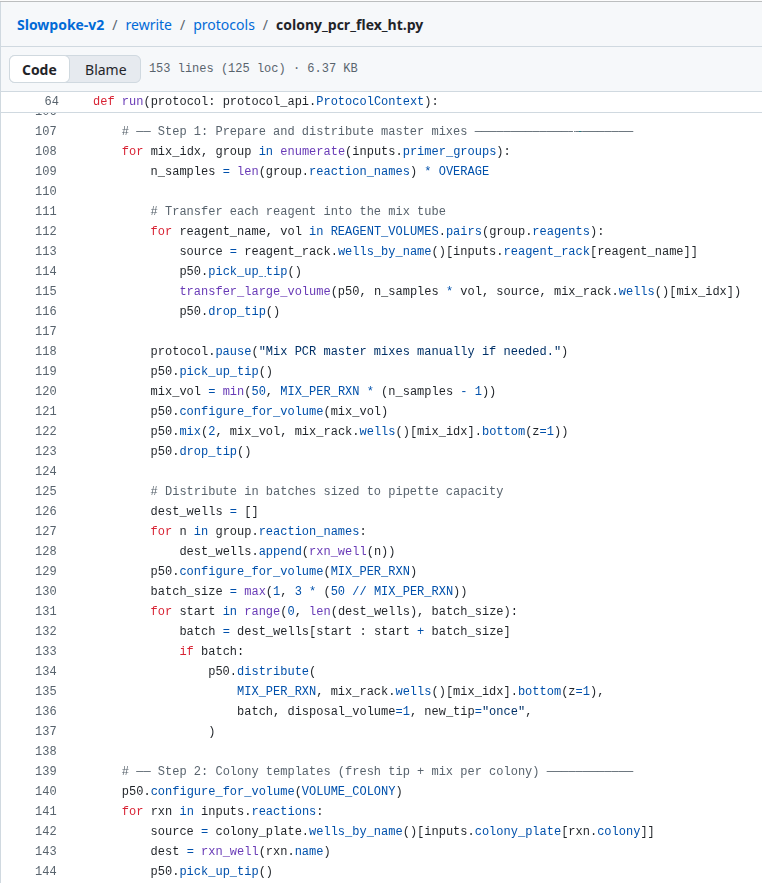
I’m not very good at following the beaten track, I always have to find a way to shape work into something I enjoy. So instead of doing the HW properly and writing some Opentrons code to paint a drawing onto agar, I decided to take a stab at rewriting the protocol the scientists published as open-source code.
The reason for doing this is part understanding, and part interest. Scientists work in a different domain to engineers, so the code is usually a very different shape to what we’re used to. I wondered what it would look like if the Opentrons protocols looked like code you would see at Google - ie. well-encapsulated, one-way flows of state, explicit control flow, good use of abstractions.
Over a couple iterations I came to this, which seemed interesting enough to finish on. It’s very interesting to read the code and see how things work. I have a few ideas from this course of what I would build as my dream bio-engineering software environment - in terms of DNA, experimentation as code, encapsulation and instrumentation. For now, I thought I’d take the chance to write about what I find interesting about the field right now.



If you find this interesting, Keoni Gandall wrote a very short book talking about this opportunity area of designing the programming environments for synthetic biology. Check it out here - https://synbio25.com/
https://github.com/liamzebedee/Slowpoke-v2/blob/main/rewrite/protocols/cloning_flex_ht.py